|
The term ‘variant’ has come into common use, even among the general public, to denote a set of such co-occurring amino acid changes 4. As the COVID-19 pandemic progressed, research interests shifted towards the study of coordinated mutations. Genomic surveillance concentrated initially on the monitoring of individual amino acid changes, such as the rise to dominance of the D614G Spike mutation worldwide 2 or the spread of the A222V Spike mutation from Spain throughout Europe during the 2020 Summer 3. Several organizations offer repositories for depositing sequences the largest collection is provided by GISAID 1, which recently hit two million deposited records. The fast emergence of SARS-CoV-2 mutations and their possibly severe epidemiological implications call for continuous and worldwide monitoring of viral genomes. Our early warning system, exclusively relying on deposited sequences, shows the power of big data in this context, and concurs to calling for the wide spreading of public SARS-CoV-2 genome sequencing for improved surveillance and control of the COVID-19 pandemic. This allows us to detect variants’ emergence, rise, peak, and eventual decline under competitive pressure of another variant. Our method succeeds in timely associating clusters to variants of interest/concern, provided their change composition is well characterized. For all countries with sufficient data, we compute weekly counts of amino acid changes, unveil time-varying clusters of changes with similar-rapidly growing-dynamics, and then follow their evolution. Here we show that the emergence of variants can in fact be traced through data-driven methods, further capitalizing on the value of large collections of SARS-CoV-2 sequences. Yet, never has this problem been tackled by digging into data with ad hoc analysis techniques. The co-occurrence of specific amino acid changes, collectively named ‘virus variant’, requires scrutiny (as variants may hugely impact the agent’s transmission, pathogenesis, or antigenicity) variant evolution is studied using phylogenetics. 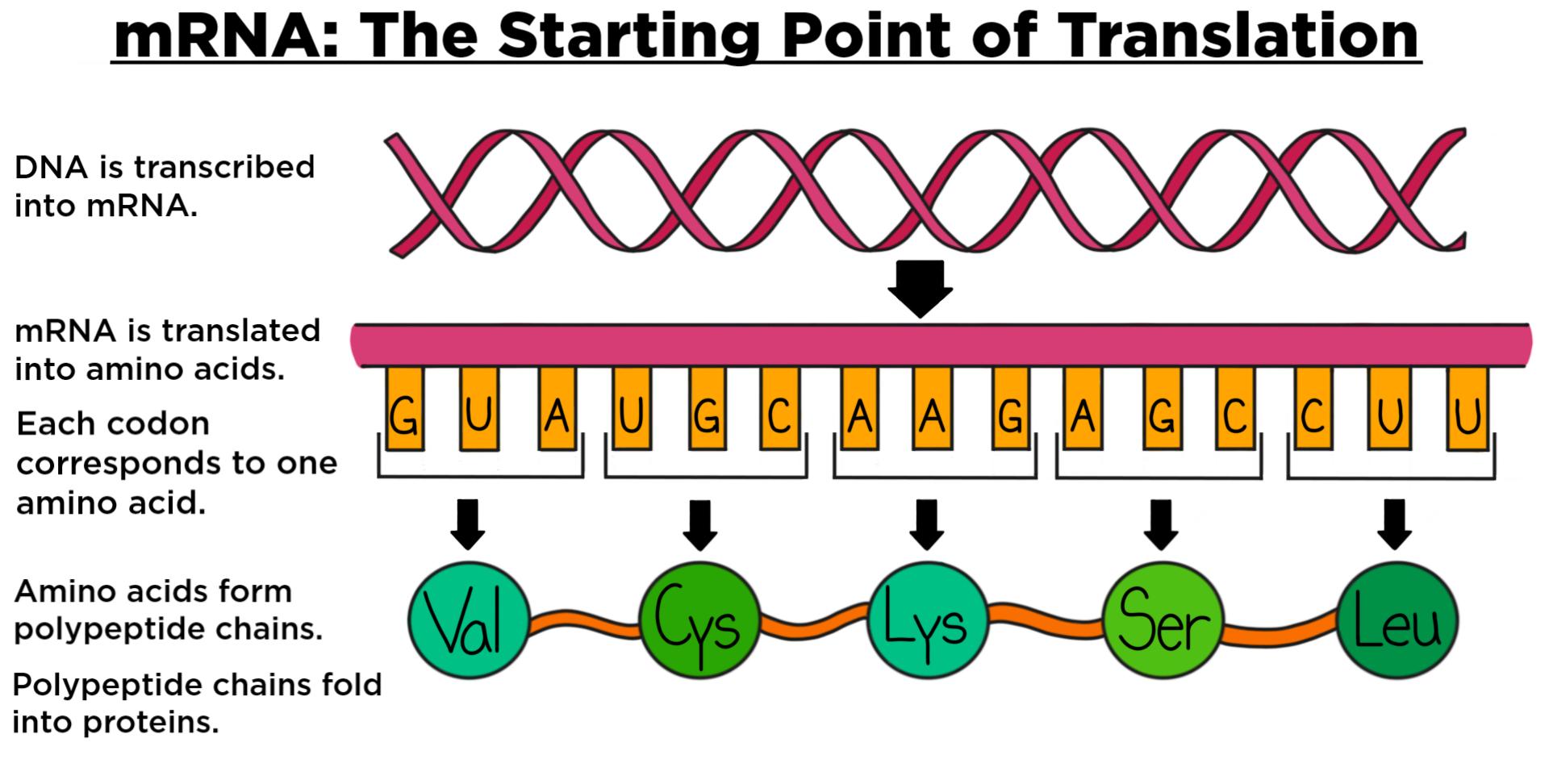 
Since its emergence in late 2019, the diffusion of SARS-CoV-2 is associated with the evolution of its viral genome.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
